
Twist Bioscience Variant Libraries | Site Saturation Variant Libraries
Generate 99% of Desired Variants
Protein engineering screens using single site variant libraries allow researchers to explore a protein’s sequence space and investigate the relationship between sequence and protein structure and function.
Twist Site Saturation Variant Library construction leverages massively parallel oligonucleotide synthesis using Twist's proprietary silicon-based DNA synthesis platform. Twist libraries are NGS-verified to confirm that all desired variants are present in the correct ratios.
KEY BENEFITS
- Complete control over codon usage (all 64 codons available)
- High uniformity of variant representation at every site
- No unwanted codons or premature stop codons
- Rigorous quality control
- Verified uniformity of variant representation by NGS
- Representation of each site normalized by mass
- One position per well in a 96 well plate or all positions pooled in a single tube
- Screen 1 to 20 different amino acids at each position
WATCH WEBINAR
Learn more about how Twist Single Site Variant Libraries (SSVL) enable the interrogation of sequence space to identify key residues in protein structure and function.
Precision Synthesis of Variant Libraries Enables Comprehensive Interrogation of Single-Site
Explore the Sequence Space
Site Saturation Libraries eliminate codon bias and unwanted substitutions unlike traditional methods such as degenerate and NNK approaches.
TRIM offers a poor repetitive yield, leading to <50% full-length product in a typical library whereas, Twist libraries yield more usable variants, increasing effective library size.
| ERROR PRONE PCR | DEGENERATE
(NNK/NNS) |
TWIST SITE SATURATION VARIANT LIBRARIES |
|
|---|---|---|---|
| Eliminates sequence bias | No | No | Yes |
| Number of codons available | Unknown | 32 | All 64 |
| Prevents undesirable motifs | No | No | Yes |
| Allows codon optimization | No | No | Yes |
| Avoids stop codons | No | Yes | Yes |
Highly Uniform Variant Libraries
Site Saturation Variant Libraries enable efficient sampling of a protein’s sequence space in screening assays.
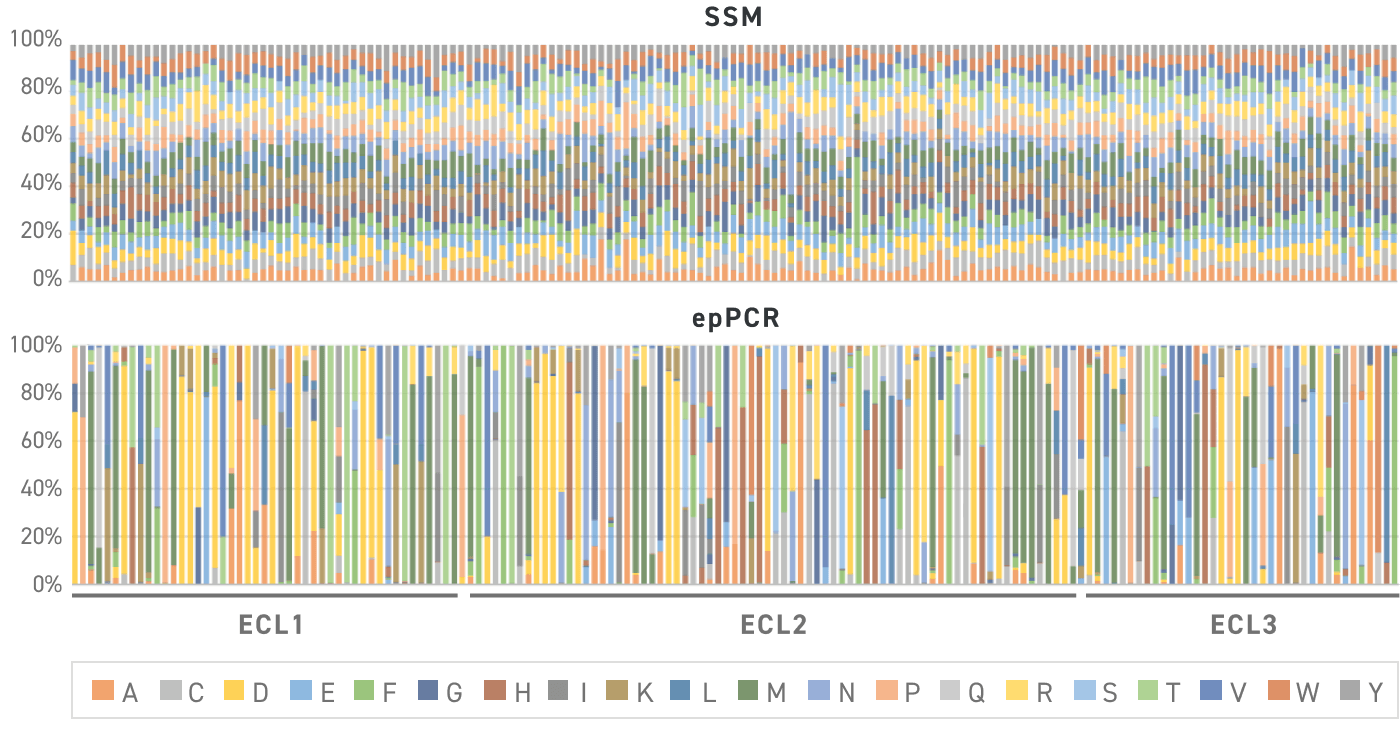
This figure shows the observed distribution of amino acids across 65 positions in protein (19 intended variants per position) when built with Twist SSVLs versus error prone PCR. The bars represent a different amino acid position, with each color indicating the observed variant frequency. All variants are present in the expected ratios in the Twist Library.
DOWNLOADS
- Precision Synthesis of variant Libraries Enables Deep Interrogation of Single-Site Variant Space - Download

